Welcome!
The Eaton lab is in the Department of Ecology, Evolution, and Environmental Biology at Columbia University. Here you can find information about news and updates in the lab, current and former members, as well as fun posts about fieldwork, discussions of papers, and new methods being developed and used in the lab. Below you can find descriptions of several major ongoing projects.
Evolutionary consequences of interspecific reproductive interactions
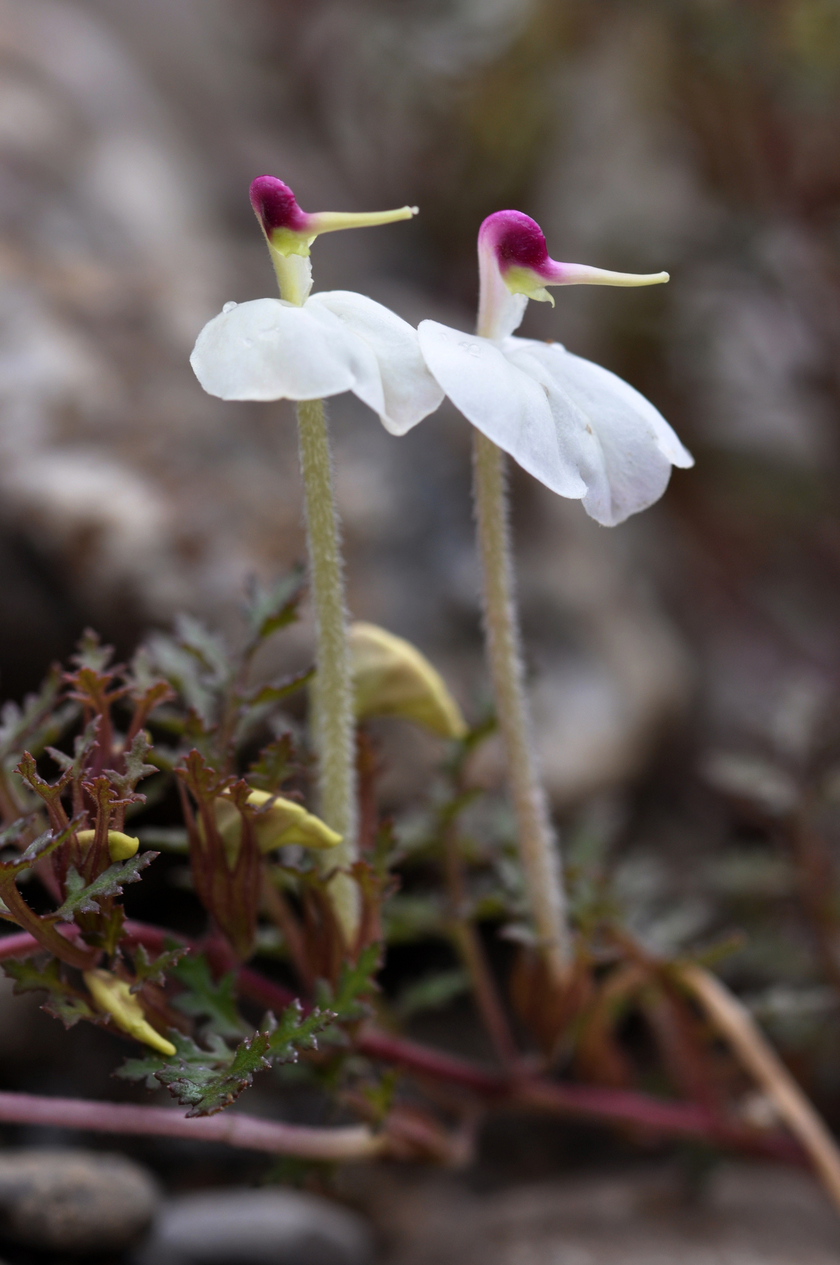
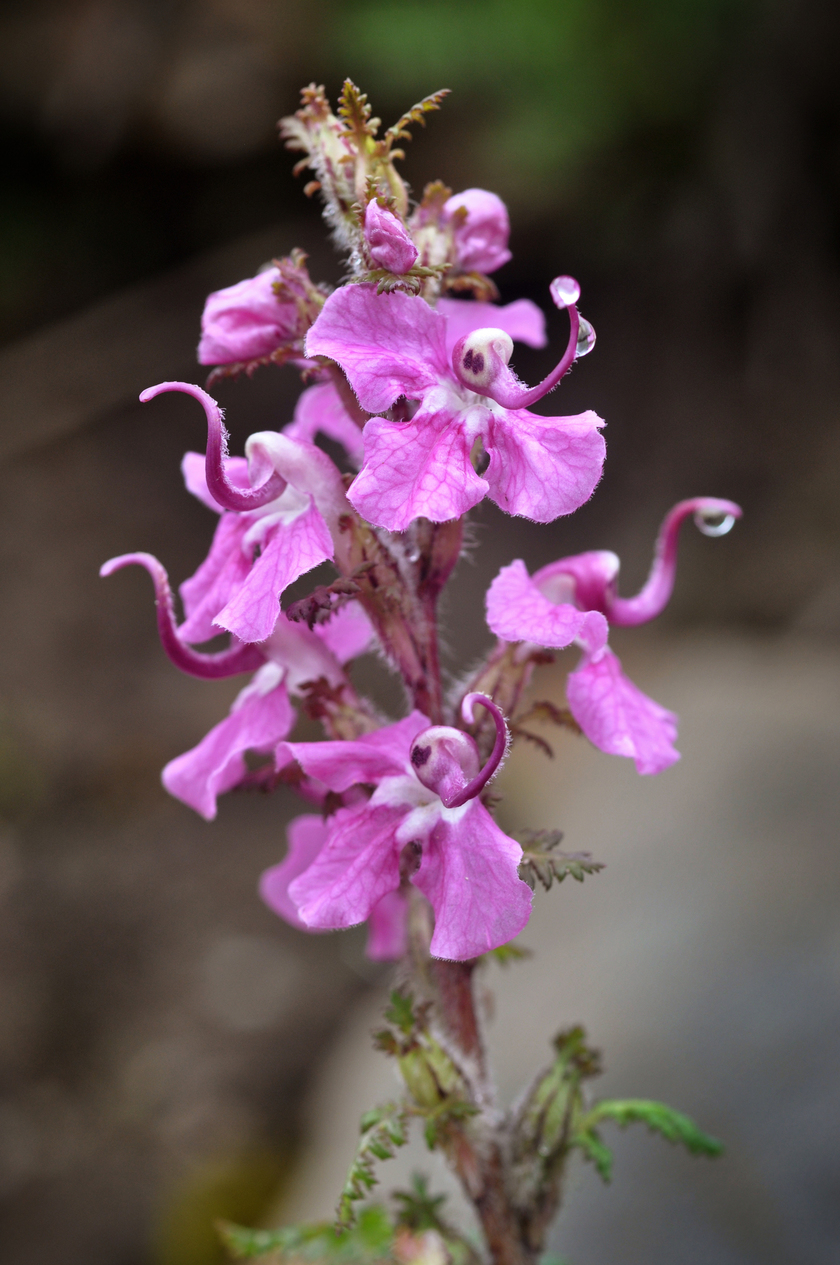
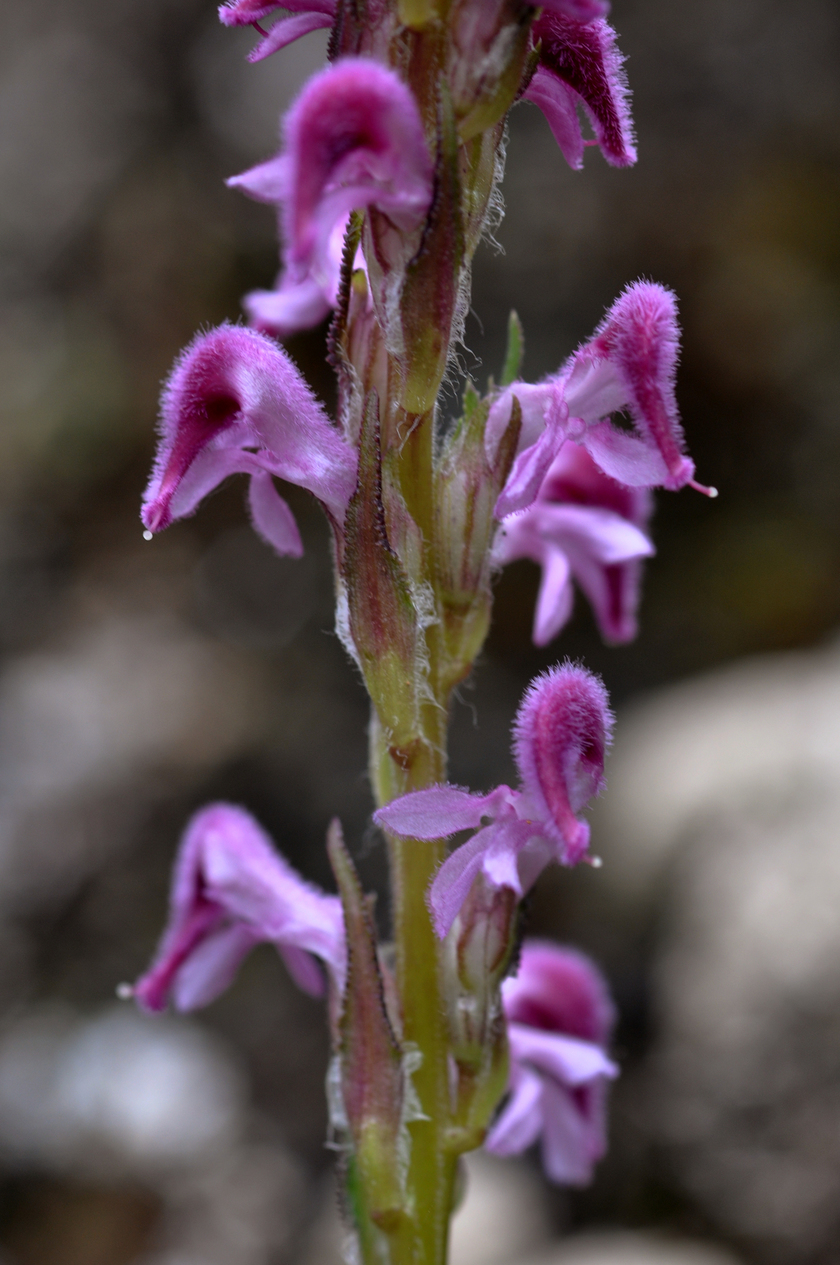
The Hengduan Mountains of China on the eastern edge of the Tibetan plateau is one of the most diverse temperate plant communities in the world. Of the many clades that have diversified in situ in this region Pedicularis stands out for its exceptional diversity of floral forms displayed across >300 endemic species. A general link between floral diversity and species diversity has long been recognized within flowering plants, and the concentrated diversity of Pedicularis in the Hengduan Mountains provides an ideal system in which to investigate the role that interspecific reproductive interactions play in shaping this pattern. The Eaton lab combines genomic analyses with field-based experiments and community analyses to understand the role that gene flow and reproductive interference play in shaping Pedicularis evolution.



Replicated evolution of leaf form in Viburnum
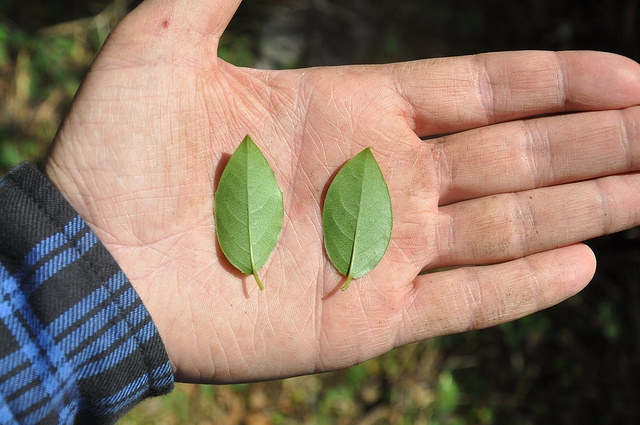
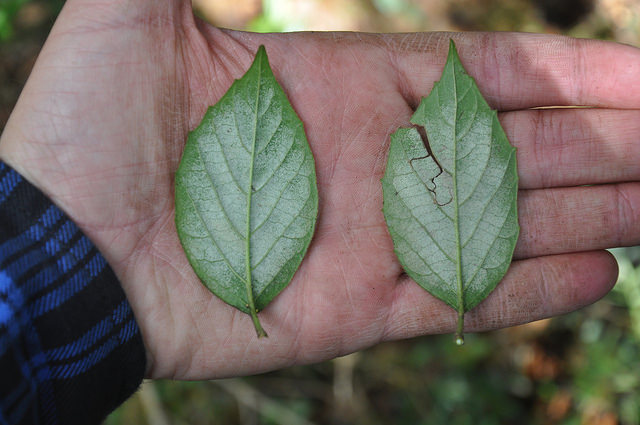
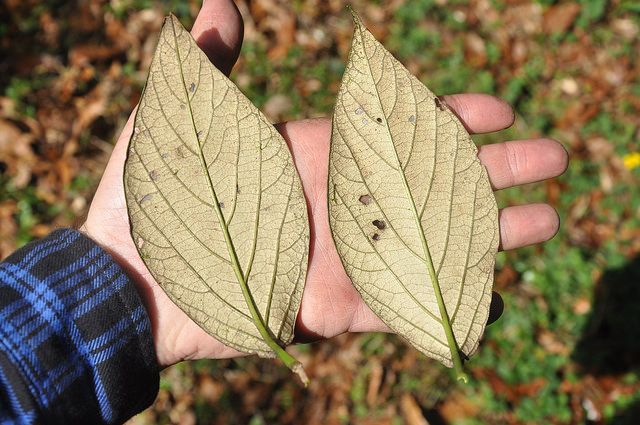
Viburnum is a widespread group of small trees and shrubs distributed primarily throughout temperate regions of the northern hemisphere. The most rapid radiation among extant Viburnum occurred in a clade that diversified throughout neo-tropical cloud forests in the last 10My. In these narrow, isolated environments stretching from Mexico to Bolivia, a distinct set of morpho-species occurs repeatedly, representing distinct sets of co-varying leaf traits. Phylogenomic analyses suggest that species within each geographical region are each others closest relatives, but hybridization also makes their relationships highly reticulate. We are combining genomic analyses with eco-physiological studies to understand species relationships and the processes underlying this apparent pattern of replicated evolution.



Computational phylogenetics and bioinformatics
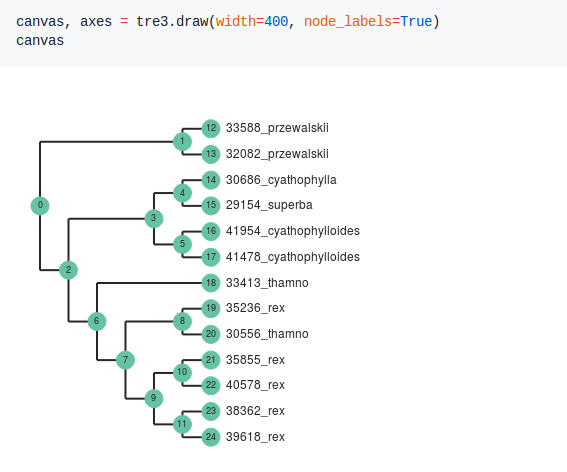
I have been writing software primarily in the Python programming language for a number of years now. This includes software for assembly of genomic RAD-seq type data sets (ipyrad), for quartet species tree inference (tetrad), and for tree plotting (toytree). A major focus of software development in the Eaton lab is to create methods that make it easy to write and share code for reproducible analyses, which when writing in Python has steered development strongly towards the use of jupyter notebooks.
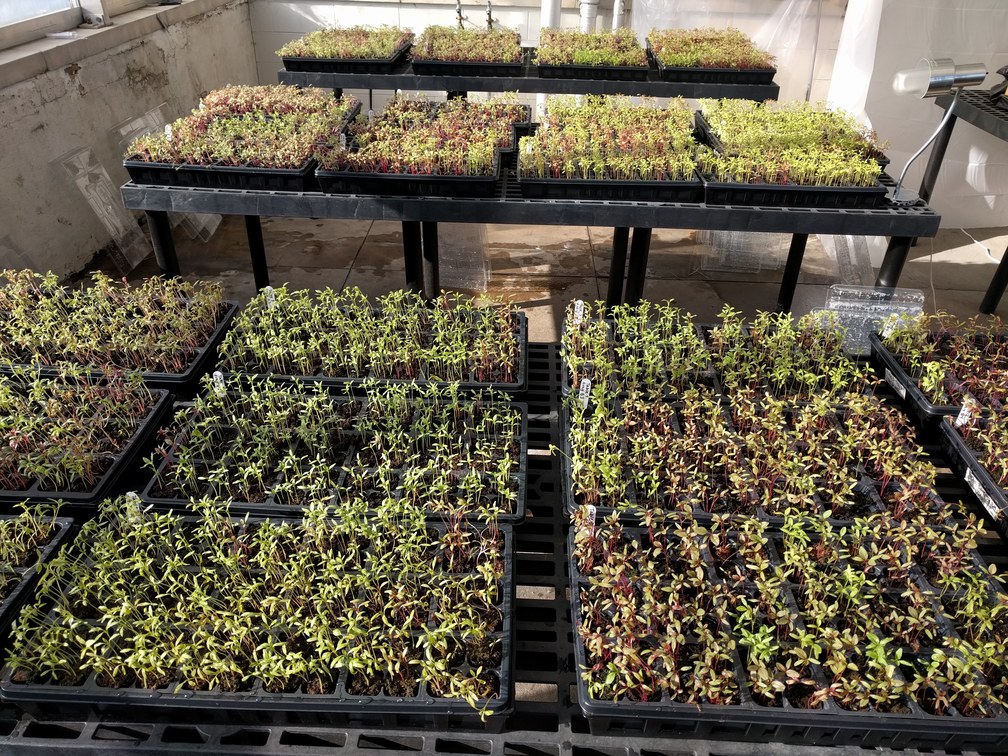
Speciation genomics in Amaranthus
The cultivated species of grain amaranth make up only a small proportion of the much larger Amaranthus clade. While a lot of work is being done to study the genome of the cultivated species, we are interested in applying these powerful genomics resources for understanding the evolutionary history of the entire Amaranthus genus. We are studying diversification, speciation, biogeography, floral evolution, and hybridization. Amaranthus is a nice model system for such studies owing to it’s fast growth, small genome size, and variable mating system strategies.

