Principles and Applications of Modern DNA Sequencing
EEEB GU4055
Session 13: Phylogenetics II
Today's topics
1. Review notebook assignments: RAD-seq
2. Discuss the assigned readings: RAD & Oaks study
3. Introduce new topic: species trees & midterm
Review of course topics
1. DNA sequencing review; and intro to Jupyter/Python.
2. Python bootcamp I: Basic objects.
3. Python bootcamp II: Scientific libraries.
4. Homology/Blast/GFF: Genome structure
5. Phylogenetics I: Sanger sequences to trees.
6. Recombination and Meiosis.
7. Inheritance and pedigrees.
8. Intro to Illumina and read mapping.
9. Intro to long-read technologies and read mapping.
10. Genome Assembly in theory.
11. Genome Assembly in practice.
12. Long-reads and Hi-C Scaffolding.
13. Phylogenetics II: RAD-seq
14. Phylogenetics II: SNPs, gene trees and species trees
Notebook 13.1: RAD-seq
Restriction-site associated DNA sequencing (RAD-seq) is a method
for subsampling the genome adjacent to restriction enzyme recognition
sites.
Provides a cheap, easy and fast method for genotyping many markers
(typically thousands) that are spread throughout the entire genome.
Notebook 13.1: RAD-seq
When a genome is digested with a restriction enzyme the genome is broken into smaller fragments. Each fragment will begin and end with a characteristic overhang. Your notebook this week emulated this process bioinformatically.
Phylogenomics II
Genomics methods allow us to efficiently sample variation across the genome.
Phylogenomics II
Background on the method and application of RAD-seq:

Phylogenomics II
Applied phylogenomics example:

Notebook 13.1: RAD-seq
Genomic library preparation for Illumina sequencing involves combining oligonucleotide molecules in solution, in a particular order, to build larger molecules that will bind by complementarity to the sequencing machine where they can be read using "sequencing by synthesis".
We built oligonucleotides bioinformatically for a RAD-seq library following the Original RAD method (*sensu* Andrews et al.) using the `PstI` restriction enzyme (CTGCA^G), and ligation of six-base barcodes (e.g., AACCTT) and the two Illumina adapters.
Notebook 13.1: RAD-seq variants: original RAD
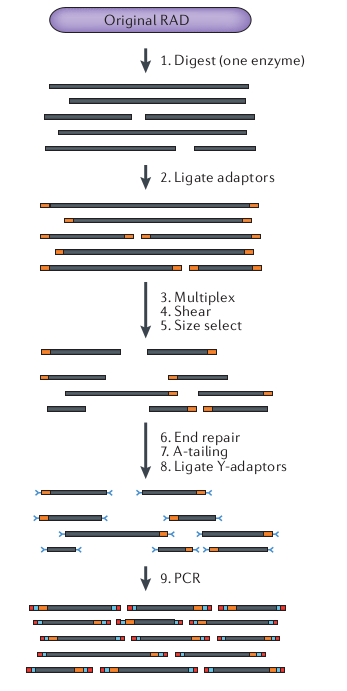
Notebook 13.1: RAD-seq variants: ddRAD
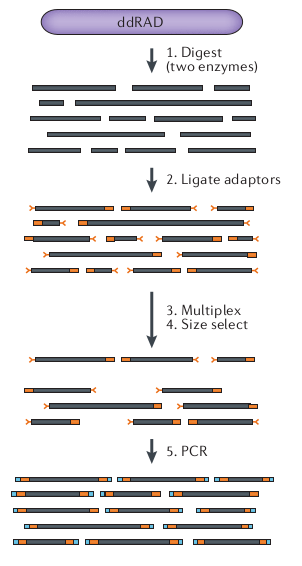
Notebook 13.1: RAD-seq
def random_sequence(length):
"return a random sequence of DNA"
return "".join(np.random.choice(list("ACGT"), size=length))
def restriction_digest(sequence, recognition, cut):
"""
restriction digest a genome sequence at the given (recognition) site and
split the site at the given position (cut) to leave overhangs. Returns one
strand of the resulting fragment: e.g., for recognition=CTGCAG, cut=5 this
returns the 5' to 3' strand below.
5' ----G[seq]CTGCA 3'
3' ACGTC[seq]G---- 5'
"""
# cut sequence at every occurence of recognition site
fragments = sequence.split(recognition)
# add overhang that results from sequence splitting within the recognition site
fragments = [recognition[cut:] + i + recognition[:cut] for i in fragments]
return fragments
Notebook 13.1: RAD-seq
def complement(sequence):
"return the complement of a sequence"
return (sequence
.replace("C", "g")
.replace("G", "c")
.replace("T", "a")
.replace("A", "t")
.upper())
Notebook 13.1: RAD-seq
def ligate_barcoded_adapters(fragments, recognition, cut, barcode):
"""
Ligates a barcoded Illumina adapter to each fragment if it has a cleaved
sequence overhang matching the recognition site. This will fill the molecule
so that both ends have full recognition sequence and Illumina adapter.
"""
ligated = []
for fragment in fragments:
# adapter has the recognition overhang complement attached to it so
# it can bind to 5' strand and complement with 3' strand.
# ----G[seq]CTGCA <- attaches left side of here
# ACGTC[seq]G----
ligate_to_this_strand = "{}{}{}".format(
complement(ILLUMINA_ADAPTER_1)[::-1],
complement(barcode)[::-1],
recognition[-cut:-(len(recognition) - cut)],
)
# attaches to this strand and complements overhang on other strand.
# ----G[seq]CTGCA <- attaches here
# ACGTC[seq]G----
ligate_to_other_strand = "{}{}".format(
barcode,
ILLUMINA_ADAPTER_1,
)
# the full molecule has the fragment in the middle
molecule = "{}{}{}".format(
ligate_to_this_strand,
fragment,
ligate_to_other_strand,
)
ligated.append(molecule)
return ligated
Notebook 13.1: RAD-seq
# generate a random genome sequence
scaffold = random_sequence(1000000)
# cut up the genome with a restriction enzyme
fragments = restriction_digest(scaffold, "CTGCAG", 5)
# attach a barcode (AATTCC) and adapter to the overhanging end sequence (CTGCA)
barcoded_fragments = ligate_barcoded_adapters(fragments, "CTGCAG", 5, "AATTCC")
How many fragments are in the library after the restriction digestion? Calculate it from the fragments or barcoded_fragments lists. If we used a restriction enzyme with a longer recognition sequence would it produce more or fewer fragments?
len(barcoded_fragments)
# 231
There are 231 fragments in the digested library. If we used a restriction enzyme that cuts at a longer recognition site is would cut less often and we would get fewer fragments.
# select first fragmented molecule
fragment = barcoded_fragments[0]
# print features of the molecule
print("{:<\40}adapter I (revcomp)".format(fragment[:33]))
print("{:<\40}barcode (revcomp)".format(fragment[33:39]))
print("{:<\40}restriction overhang (revcomp)".format(fragment[39:44]))
print("{}... 5' end of strand".format(fragment[44:74]))
print("...{} 3' end of strand".format(fragment[-74:-44]))
print("{:<\40}restriction overhang".format(fragment[-44:-39]))
print("{:<\40}barcode".format(fragment[-39:-33]))
print("{:<\40}adapter I".format(fragment[-33:]))
TGACTGGAGTTCAGACGTGTGCTCTTCCGATCT adapter I (revcomp)
GGAATT barcode (revcomp)
TGCAG restriction overhang (revcomp)
TCTCGTCAGCTGGACGCAACAGTGTTATCT... 5p end of strand
...CCAGGATAACCGTAATTCAATTGCAGTCAC 3p end of strand
CTGCA restriction overhang
AATTCC barcode
AGATCGGAAGAGCACACGTCTGAACTCCAGTCA adapter I
def shear_DNA_to_size_select(fragments, size):
"randomly shears fragments down to the entered size"
sheared_fragments = []
# iterate over fragments
for fragment in fragments:
# get breakpoints for shearing into chunks of size 'size'
nfrags = max(2, int(len(fragment) / size))
splits = np.linspace(0, len(fragment), nfrags).astype(int)
# slice out bits to represent sheared fragments
idx = 0
for end in splits[1:]:
fragbit = fragment[idx:end]
sheared_fragments.append(fragbit)
idx = end
return sheared_fragments
# break into 300bp bits and only keep bits
sheared_fragments = shear_DNA_to_size_select(all_fragments, 300)
How many fragments are in the library after shearing? What type of sequences do you expect to find at the ends of the fragments after shearing? How is this different from the expectation after the initial restriction digestion? Answer in Markdown below.
len(sheared_fragments)
# 12176
After shearing there are 12176 fragments in the library since many of the fragments were longer than 300 bp and we sheared them all down to this size. The ends of the sheared fragments are random positions, but some will still have a restriction overhang on one end. This is different from before shearing, when each fragment had a restriction overhang on each end.
Question: Look at the contents of the fastq files in the following directory: /home/jovyan/ro-data/SRP021469/ Based on the sequence data in each file, and what you now know about the structure of oligonucleotides that make up RAD-seq data, what was the restriction enzyme that was used to build this RAD-seq library? For the file 29154_superba_SRR1754715.fastq.gz in this directory, what is the barcode sequence? Hint: it may help to compare the sequences found in several files (data from several different individuals) to figure out which is the barcode sequence and which is the restriction sequence. Answer in Markdown below.
In the sentence above this question I suggest 2 ways: (1) Use the unix tools `less` or `head` to peek at the contents of the file; or (2) use Python to load the file and print the first few lines. We've learned both of these things before.
import gzip
data = gzip.open("/home/jovyan/ro-data/SRP021469/29154_superba_SRR1754715.fastq.gz")
# print a few lines
for i in range(10):
line = data.readline().strip().decode()
print(line)
data.close()
@29154_superba_SRR1754715.1 GRC13_0027_FC:4:1:12560:1179 length=74
TGCAGGAAGGAGATTTTCGNACGTAGTGNNNNNNNNNNNNNNGCCNTGGATNNANNNGTGTGCGTGAAGAANAN
+29154_superba_SRR1754715.1 GRC13_0027_FC:4:1:12560:1179 length=74
IIIIIIIGIIIIIIFFFFF#EEFE<\?###############################################
@29154_superba_SRR1754715.2 GRC13_0027_FC:4:1:15976:1183 length=74
TGCAGTTGTAAATACAAATATCCCAAAANNNNGNNNNNNNTNTAATATTTTGNAANNTTGAGGGGTGTGATNTN
+29154_superba_SRR1754715.2 GRC13_0027_FC:4:1:15976:1183 length=74
GGGGHHHHHHHHHHHHHDHGHHHHCAAA##############################################
@29154_superba_SRR1754715.3 GRC13_0027_FC:4:1:19092:1179 length=74
TGCAGGCTCTGACAAAGAANTCGACTGANNNNNNNNNNNNNNCACNGGTTCNNGNNNATGTCAATGTGGTANAN
Or, open a terminal (from the dashboard page, as explained in the instructions) and use the `less` or `zcat [file] | head` commands.
# opens an interactive view to one window full of the file. (q to quit).
less /home/jovyan/ro-data/SRP021469/29154_superba_SRR1754715.fastq.gz
# or, print first n lines to the screen
zcat /home/jovyan/ro-data/SRP021469/29154_superba_SRR1754715.fastq.gz | head -n 20
Notebook 13.2: RAD-seq pipelines
The ipyrad software is used to make variant calls and assemble loci from
RAD-seq reads. Two approaches: de novo and reference. Reference mapped data
uses the mapping coordinates to identify orthology of sequences across samples.
Without a reference clustering is used to identify orthology based on
pairwise sequence similarity.
ipyrad is convenient to use as it provides a full pipeline from raw data to
assembled loci, and includes many analysis tools specialized for RAD data.
Notebook 13.2: Object oriented Python library
import ipyrad as ip
# create an Assembly object
data = ip.Assembly("test")
# view default params of object
data.params
# set parameters for assembly
data.params.raw_fastq_path = "/home/jovyan/ro-data/ipsimdata/rad_example_R1_.fastq.gz"
data.params.barcodes_path = "/home/jovyan/ro-data/ipsimdata/rad_example_barcodes.txt"
...
# run steps of the assembly
data.run("123", auto=True)
... 
Notebook 13.2: After assembly
# final assembled data files can be accessed from the object directly
data.outfiles
import ipyrad.analysis as ipa
import toytree
# infer a ML tree
rax = ipa.raxml(data=data.outfiles.phy, N=10)
rax.run(block=True, force=True)
# plot the tree
tre = toytree.tree(rax.trees.bipartitions)
tre.root(wildcard="3").draw();
Look at the output files (again, I recommend using `less`).
# opens an interactive view to one window full of the file. (q to quit).
less /home/jovyan/work/simdata/test_outfiles/test.loci
reference TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
1A_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
1B_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
1C_0 TTTAACTGTTCAAGTCGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
1D_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
2E_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
2F_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
2G_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
2H_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCGAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
3I_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
3J_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
3K_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCCC
3L_0 TTTAACTGTTCAAGTTGGCAAGATCAAGTCGTCCCTAGCCCCCGCGTCCGTTTTTACCTGGTCGCGGTCCCGACCCAGCTGCCCSC
// - - - |0:MT:5014-5100|
reference GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
1A_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
1B_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
1C_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGATCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
1D_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
2E_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
2F_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
2G_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
2H_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
3I_0 GCGACTCTATGGAGGAAGGCWCACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
3J_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
3K_0 GCGACTCTATGGAGGAAGGCACACCCGCCATTGCAGGTCATCAACTATACTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
3L_0 GCGACTCTTTGGAGGAAGGCACACCCGCCATTGCAGGTCATCAATTATAGTGAGTGCCGTGTTGGCACCATCCAGTGTGAATTACA
// - - - - - |1:MT:10114-10200|
reference CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
1A_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
1B_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
1C_0 CACTAGGACCCCCATATAGGKTATAGGGTWGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCRCGTCCAGCTTAATTAC
1D_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATCAC
2E_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
2F_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
2G_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
2H_0 CACTAGGACCCCCATATAGGTAATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCRTCCAGCTTAATTAC
3I_0 CACTAGAACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
3J_0 CACTAGRACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
3K_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
3L_0 CACTAGGACCCCCATATAGGTTATAGGGTTGCTATGTATAAGCCGTCGGCAAATTCCGCGAACGCTGTCGCGTCCAGCTTAATTAC
// * -- - - - - |2:MT:15214-15300|
Let's look at an empirical example:
# opens an interactive view to one window full of the file. (q to quit).
less /home/jovyan/ro-data/SRP021469_outfiles/SRP021469.loci
29154_superba_SRR1754715 TCATGGATTTTATTAATTTTG--TAAAAAAGATTGAATAAGTATGTTCAATAGAATATATTATGCTTGAC
30556_thamno_SRR1754720 TCATGGATTTTATTAATTTTG-AAAAAAAAGATTGAATAAGTATGTTTAATATAATATATTATGCTTGAC
30686_cyathophylla_SRR1754730 TCATGGATTTTATTAATTTTG--TAAAAAAGATTGAATAAGTATGTTCAATAGAATATATTATGCTTGAC
33413_thamno_SRR1754728 TCATGGATTTTATTAATTTTG--TAAAAAAGATTGAATAAGTATGTTTAATATAATATATTATGCTTGAC
35236_rex_SRR1754731 TCATGGATTTTATTAATTTTG-AAAAAAAAGATTGAATAAGTATGTTTAATATAATATATTATGCTTGAC
35855_rex_SRR1754726 TCATGGATTTTATTAATTTTG-AAAAAAAAGATTGAATAAGTATGTTTAATATAATATATTATGCTTGAC
40578_rex_SRR1754724 TCATGGATTTTATTAATTTTGAAAAAAAAAGATTGAATAAGTATGTTTAATATAATATATTATGCTTCAC
// * * * - |0|
29154_superba_SRR1754715 GGNGAATATTGTTTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
30556_thamno_SRR1754720 GGTGAATATTGTTTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
30686_cyathophylla_SRR1754730 GGNGAATATTGTTTTCCACGACAATGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
33413_thamno_SRR1754728 GGTGAATATTGTTTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
35236_rex_SRR1754731 GGTGAATATTGTTTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
35855_rex_SRR1754726 GGTGAATATTGTTTTCCATGACATTGATAAAACTGATCATCTTTAAGAACAACCCAAGACATGATCAAA
38362_rex_SRR1754725 GGTGAATATTGTGTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
39618_rex_SRR1754723 GGTGAATATTGTGTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
40578_rex_SRR1754724 GGTGAATATTGTTTTCCAYGACATTGATAAAACTGATCATCTTTAAGAACWACCCAAGACATGATCAAA
41478_cyathophylloides_SRR1754722 GGTGAATATTGTTTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
41954_cyathophylloides_SRR1754721 GGTGAATATTGTTTTCCACGACATTGATAAAACTGATCATCTTTAAGAACTACCCAAGACATGATCAAA
// * * - * |1|
29154_superba_SRR1754715 AACACCTGAAATAAATTTTATGTGCAATGCACAC--ACACACACCTCCCATGCGTGTGTGTCAGAAANA
30556_thamno_SRR1754720 AACACCTGAAATAAATTTTATGTGCAACGCACACAAACACACACCTACCATGCGTGTGTGTCAGAAAAA
30686_cyathophylla_SRR1754730 AACACCTGAAATAAATTTTATGTGCAATGCACACACACACACACCTCCCATGCGTGTGTGTCAGAAAAA
35236_rex_SRR1754731 AACACCTGAAATAAATTTTATGTGCAATGCACACAAACACACACCTCCCATGCGTGTGTGTCAGAAAAA
35855_rex_SRR1754726 AACACCTGAAATAAATTTTATGTGCAATGCACACAAACACACACCTCCTATGCATGTGTGTCAGAAAAA
38362_rex_SRR1754725 AACACCTGAAATAAATTTTATGTGCAATGCACACAAACACACACCTCCCATGCGTGTGTGTCAGAAAAA
39618_rex_SRR1754723 AACACCTGAAATAAATTTTATGTGCAATGCACACAAACACACACCTCCCATGCGTGTGTGTCAGAAAAA
40578_rex_SRR1754724 AACACCTGAAATAAATTTTATGTGCAATGCACACAAATACACACCTCCCATGCGTGTGTGTCAGAAAAA
// - - - - - - |2|
Phylogenomics II
Inference of phylogeny and introgression from multi-locus genomic data.
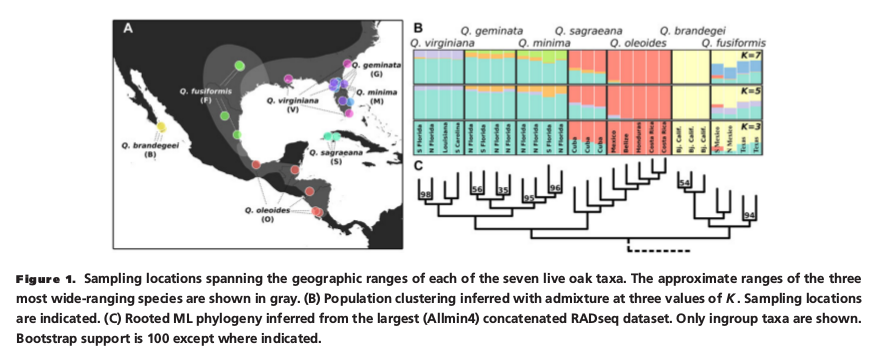
See Table 1 in the Eaton et al. paper which shows the number of RAD-seq loci that were included in de novo assembled data sets that were assembled with different parameter settings. The Allmin4 and Allmin20 assemblies differ greatly in the total number of loci, the number of informative sites, and the missing data. Why do you think?
RAD-seq data is characterized by missing data because many loci will be sequenced for some samples but not for others. This can occur from mutations to restriction sites, from low sequencing coverage, or from the generation of new restriction sites in only a subset of samples. These data sets were assembled with different parameters to yeild data sets with different amounts of missingness. Lots of missing data is not a problem for phylogenetic inference. It is a big problem for many population genetic methods like principal components analysis. In this paper Eaton et al. found missingness had no effect on phylogeny, and they used a data set with very little missing data (allmin20) for structure analyses.
Two important considerations for inferring introgression:
(1) Missing samples (ghost lineages) may be the true source of introgression.
(2) Phylogenetic relatedness (non-independence) can give a spurious signal of
introgression.
Phylogenomics II: the coalescent
Genealogical relationships vary across the genome because the genome represents a mosaic of chunks inherited from many different ancestors. This variation can be modeled using the coalescent -- a mathematical approximation for the relationship between population size and the expected number of generations back in time until two individuals share a common ancestor.